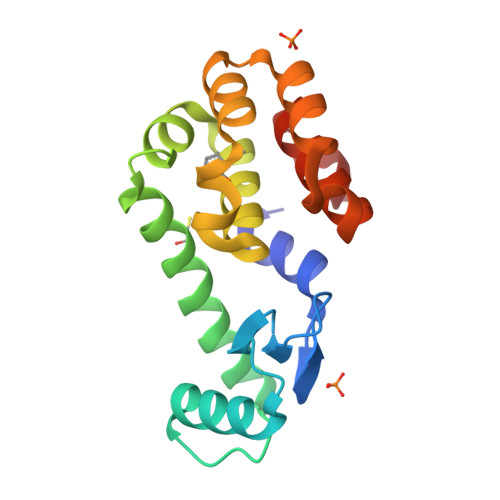
Docking
Docking
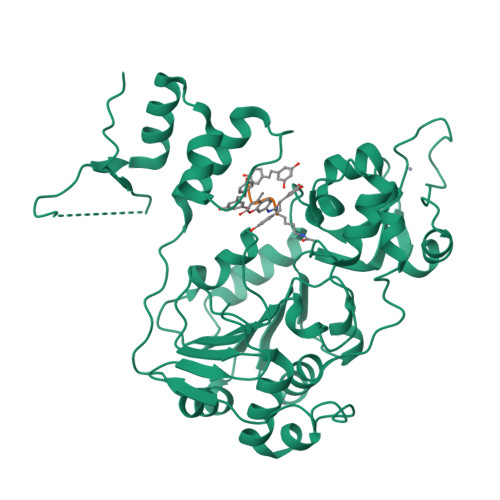 NP target
NP target
 Target
Target
 Anti-inflammatory
Anti-inflammatory
Where a molecule fits determines what it cures.
Bionics AI computes high-precision molecular docking at the binding pocket level. Given a protein structure and a natural compound SMILES, the platform identifies optimal binding poses, scores free energy of binding, and ranks candidates by estimated affinity — with full residue-level interaction maps.
Structures are energy-minimized via molecular dynamics before docking, ensuring pocket geometry reflects biologically realistic conditions, not idealized templates.